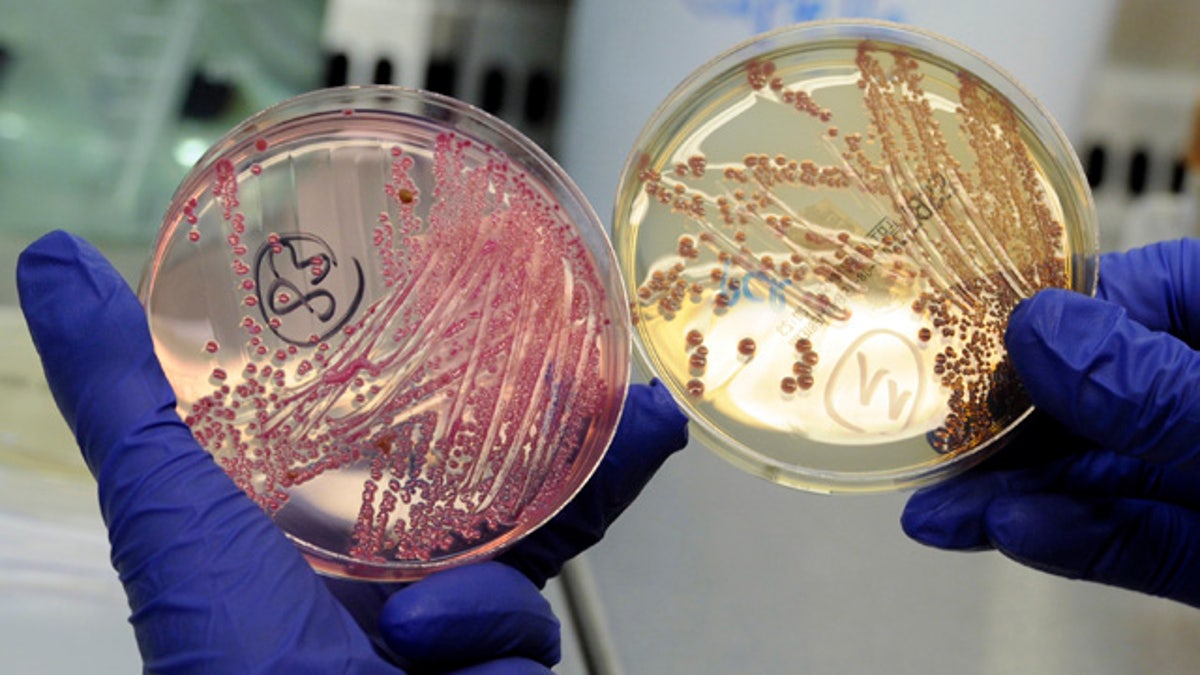
An employee holds petri dishes with bacterial strains of EHEC bacteria (bacterium Escherichia coli) in the microbiological laboratory of the Universitaetsklinikum Hamburg-Eppendorf (University Clinic Eppendorf- UKE) in the northern German town of Hamburg, June 2, 2011. (Reuters)
When most think of E. coli, they think of food poisoning, but what if the bacteria could be the key to fighting antibiotic resistance?
Researchers at the University of Buffalo in Buffalo, New York, have engineered Escherichia coli (E. coli) to produce dozens of new forms of a popular antibiotic, three of which have the ability to kill antibiotic-resistant bacteria.
Dr. Blaine Pfeifer, lead study author and associate professor of chemical and biological engineering at the University at Buffalo School of Engineering and Applied Sciences, has been working on the study for over a decade. The result is new forms of the popular antibiotic erythromycin that have a different structure from existing versions.
Pfeifer told FoxNews.com that using E. coli is a surprisingly great way to engineer erythromycin.
“One of our primary reasons for choosing to work with E. coli in the first place is that it provides numerous technical advantages including a rapid growth rate, ease of genetic manipulation, scalability for process development and production and safety. The strains we work with are all safe,” he said.
To engineer the new drugs, researchers transplanted more than 20 enzymes into E. coli and combined chemical compounds called metabolic precursors in an assembly-line process to produce the original erythromycin compound. Then, they applied the new production platform to the flexible biosynthetic pathway of erythromycin. The combination resulted in a variety of new analogs. Three of the new varieties proved to successfully kill the bacteria Bacillus subtilis, which was resistant to the original form of erythromycin.
Erythromycin is used to treat certain infections caused by bacteria like bronchitis, whooping cough, pneumonia and ear, urinary tract and skin infections. It is also used before some surgery or dental work to prevent infection.
With antibiotic resistance on the rise, and a hot topic in the world of medicine, Pfeifer said the study is especially relevant.
“There’s still a series of experiments and tests that must be completed before a final new drug is available. Having said that, the goal would be to use the new analogs of erythromycin against those bacteria that have developed resistance to the original compound,” he said.
According to Pfeifer, the findings open the door for additional engineering possibilities in the future, and could lead to even more new forms of the drug.
“The analogs produced here are really just a small sub-set of those possible. So, one future direction is to further test the analog-producing limit for the platform we have at hand,” he said.
Researchers have a goal of pushing the new antibiotics through the clinical tests needed to result in an FDA-approved drug. Then, they plan to use the approach on other current antibiotics on the market in hopes of making them capable of overcoming antibiotic resistance as well.
“There are opportunities to combat the mechanisms bacteria use to develop resistance to antibiotics," Pfeifer said. “In other words, our system allows us to engage in a game of molecular chess between mechanisms bacteria use to thwart antibiotics and the engineering capabilities we have to produce new compounds.”
The findings are published in the journal Science Advances.
